About SCON
SCON (Short Conditional intrON) is a next-generation genetic tool designed to radically simplify functional studies across 300+ vertebrate species. With a compact length of only 189 bp, SCON enables the rapid generation of conditional knockout alleles through a streamlined, one-step process.
What is SCON?
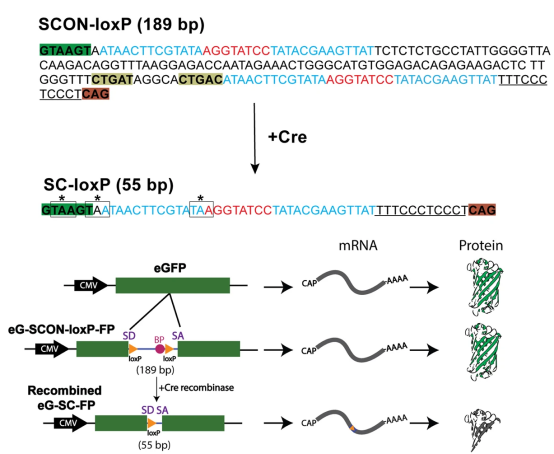
SCON is a high-performance, 189-bp artificial intron engineered for the rapid generation of conditional knockout (cKO) models across multiple species. Originally derived from the human HBB (Hemoglobin subunit beta) gene, SCON has been optimized to serve as a "genetic switch" that can be integrated into a single exon using CRISPR/Cas9-mediated knock-in.
Mechanism of Action
1. Default "ON" State:
Upon insertion into a coding exon, SCON is recognized by the cellular spliceosome as a standard intron. It is precisely spliced out during mRNA processing, leaving the protein-coding sequence intact and maintaining wild-type expression levels.
2. Conditional "OFF" State (Knockout):
The SCON sequence contains recombination sites (e.g., LoxP or FRT) flanking its critical splicing elements. When exposed to Cre or Flp recombinase, the SCON sequence is inverted or its splicing signals are disrupted. This leads to:
Intron Retention/Mis-splicing: The SCON sequence is no longer spliced out.
Premature Termination: The retained sequence introduces multiple stop codons and triggers a frame-shift, resulting in complete loss of gene function.
Key Advantages for Researchers
One-Step Zygote Injection
Due to its compact size (189 bp), SCON can be delivered via ssODN (single-stranded DNA) donors through a single injection, bypassing the need for complex and time-consuming ES cell targeting.
Universal Applicability
As demonstrated in the latest research, SCON's functionality is highly conserved across vertebrates, including mice, human iPSCs, zebrafish, Xenopus, and various species.
Minimal Hypomorphic Risk
Unlike traditional "floxing," SCON is placed within a single exon, minimizing the risk of reduced gene expression in the "ON" state.
Database Construction & Filtering
To ensure high-precision targeting and robust conditional knockout performance, our database was constructed using a systematic filtering pipeline.
1. Filtering Targetable Exons
The pipeline begins by identifying exons that are suitable for SCON insertion based on their structural and functional properties:
- •
CDS Location: Only exons located within the first 50% of the coding sequence (CDS) are selected to ensure that any induced truncation results in a complete loss-of-function.
- •
Split Exon Balance: To maintain splice donor(SD) and splice acceptor(SA) stability, the insertion site must leave at least 60 bp of exon sequence on both the 5' and 3' sides of the SCON (Left_remain & Right_remain >= 60 bp).
- •
Exon Length: A minimum length of 120 bp is required for the target exon.
2. Choose Splice Junction Motif
Following the optimized splicing consensus, sites are chosen by their flanking 4-nucleotide motifs:
- •
MAGR (Stringent): Targets (A/C)AG | (A/G) sites. This represents the most conserved mammalian splice junction for maximum splicing efficiency.
- •
VDGN (Flexible): Targets (A/G/C)(A/T/G)G | (A/T/G/C) sites, allowing for broader genomic coverage.
- •
Insertion Site: The SCON is precisely inserted between the 3rd and 4th nucleotides of the identified motif.